Abstract
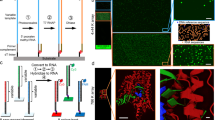
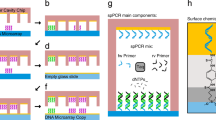
The technology and applications of microarrays of immobilized DNA or oligonucleotides are reviewed. DNA arrays are fabricated by high-speed robotics on glass or nylon substrates, for which labeled probes are used to determine complementary binding allowing massively parallel gene expression and gene discovery studies. Oligonucleotide microarrays are fabricated either by in situ light-directed combinatorial synthesis or by conventional synthesis followed by immobilization on glass substrates. Sample DNA is amplified by the polymerase chain reaction (PCR), and a fluorescent label is inserted and hybridized to the microarray. This technology has been successfully applied to the simultaneous expression of many thousands of genes and to large-scale gene discovery, as well as polymorphism screening and mapping of genomic DNA clones.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Southern, E.M. 1996. DNA chips: analysing sequence by hybridization to oligonucleotides on a large scale. Trends Genet. 12(3):110–115.
O'Donnell-Maloney, M.J., Smith, C.L., and Cantor, C.C., 1996. The development of microfabricated arrays for DNA sequencing and analysis. Trends Biotech. 14: 401–407.
Ginot, F. 1997. Oligonucleotide microarrays for identification of unknown mutations: how far from reality? Human Mutation 10: 1–10.
Drmanac, R., Drmanac, S., Labat, I., Crkvenjakov, R., Vicentric, A., and Gemmell, A. 1992. Sequencing by hybridization: towards an automated sequencing of one million M13 clones arrayed on membranes. Electrophoresis 13: 566–573.
Schena, M., Shalon, D., Davis, R.W., and Brown, P.O. 1995. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270: 467–470.
Wallraff, G., Labadie, J., Brock, P., DiPietro, R., Nguyen, T., Huynh, T., et al. 1997. DNA sequencing on a chip. Chemtech February, pp.22–32.
Schena, M. 1996. Genome analysis with gene expression microarrays. Bioessays 18: 427–431.
Shalon, D., Smith, J.S., and Brown, P.O. 1996. A DNA microarray system for analyzing complex DNA samples using two-color fluorescent probe hybridization. Genome Res. 6: 639–645.
Drmanac, S., Stavropoulos, N.A., Labat, I., Vonau, J., Hauser, B., Soares, M.B., and Drmanac, R. 1996. Gene-representing cDNA clusters defined by hybridization of 57,419 clones from infant brain libraries with short oligonucleotide probes. Genomics 37: 29–40.
Drmanac, S. and Drmanac, R. 1994. Processing of cDNA and genomic kilobase-size clones for massive screening, mapping and sequencing by hybridization. BioTechniques 17: 328–336.
Milosavljevic, A., Savkovic, S., Crkvenjakov, R., Salbego, D., Serrato, H., Kreuzer, H., et al. 1996. DNA sequence recognition by hybridization to short oligomers: experimental verification of the method on the E. coli genome. Genomics 37: 77–86.
Fodor, S.P.A., Read, L.J., Pirrung, M.C., Stryer, L., Lu, A.M., and Solas, D. 1991. Light-directed, spatially addressable parallel chemical synthesis. Science 251: 767–773.
Pease, A.N., Solas, D., Sullivan, E.J., Cronin, M.T., Holmes, C.P., and Fodor, S.P.A. 1994. Light-generated oligonucleotide arrays for rapid DNA sequence analysis. Proc. Natl. Acad. Sci. USA 91: 5022–5026.
Hacia, G.H., Brody, L.C., Chee, M.S., Fodor, S.P.A., and Collins, F.S. 1996. Detection of heterozygous mutations in BRCA1 using high-density oligonucleotide arrays and two color fluorescence analysis. Nat. Genet. 14: 441–447.
McGall, G., Labadie, J., Brook, P., Wallraff, G., Nguyen, T., and Hinsberg, W. 1996. Light-directed synthesis of high-density oligonucleotide arrays using semiconductor photoresists. Proc. Nat. Acad. Sci. USA 93: 13555–13560.
Yershov, K., Barsky, V., Belgovskiy, A., Kirillov, E., Kreindlin, E., Ivanov, I, et al. 1996. DNA analysis and diagnostics on oligonucleotide chips. Genetics 93: 4913–4918.
Heller, M.J. and Tu, E. 1993. Active programmable electronic devices for molecular biomolecules. US Pat. 5605662.
DeRisi, J.L., Iyer, V.R., and Brown, P.O. 1997. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 270: 680–686.
Schena, M., Shalon, D., Heller, R., Chai, A., Brown, P.O., and Davis, R.O. 1996. Parallel human genome analysis: microarray-based expression monitoring of 1000 genes. Proc .Nat Acad. Sci. USA 93: 10614–10619.
Heller, R.N., Schena, M., Chai, A., Shalon, D., Bedilion, T., Gilmore, J., et al. 1997. Discovery and analysis of inflammatory disease-related genes using cDNA microarrays. Proc. Natl. Acad. Sci. USA 94: 2150–2155.
DeRisi, J., Penland, L., Brown, P.O., Bittner, M.L., Meltzer, P.S., Ray, M., et al. 1996. Use of a cDNA microarray to analyse gene expression patterns in human cancer. Nat. Genet. 14: 457–460.
Iyer, V., DeRisi, J., Eisen, M., Ross, D., Spellman, P., Hudson, J. Jr., et al. 1997. Use of DNA microarrays to monitor differential gene expression in yeast and humans. Fed. Am. Soc. Exp. Bio. 11: 1126.
Wodicka, L., Dong, H., Mittmann, M., Ho, M-H., and Lockhart, D.J. 1997. Genome-wide expression monitoring in Saccharomyces cerevisiae. Bio/Technology 15: 1–15.
Lockhart, D.J., Dong, H., Byrne, M.C., Follettie, M.T., Gallo, M.V., Chee, M.S., et al. 1996. Expression monitoring by hybridization to high-density oligonucleotide arrays. Bio/Technology 14: 1675–1680.
De Saizieu, A., Certa, U., Warrington, J., Gray, C., Keck, W. and Mous, J. 1998. Bacterial transcript imaging by hybridization of total RNA to oligonucleotide arrays. Bio/Technology 16: 45–48.
Stipp, D. 1997. Gene chip breakthrough. Fortune 3/31: 56–73.
Shoemaker, D.D., Lashkari, D.A., Morris, D., Mittmann, M., and Davis, R.W. 1996. Quantitative phenotypic analysis of yeast deletion mutants using a highly parallel molecular bar-coding strategy. Nat. Genet. 14: 450–456.
Lipshutz, R.J., Morris, D., Chee, M., Hubbell, E., Kozal, M.J., Shah, N., et al. 1995. Using oligonucleotide probe arrays to access genetic diversity. Biotechniques 19: 442–447.
Kozal, M.J., Shah, N., Shen, N., Yang, R., Fucini, R., Merigan, T.C., et al. 1996. Extensive polymorphism observed in HIV-1 clade B protease gene using high-density oligonucleotide arrays. Nature Medicine 2: 753–759.
Cronin, M.T., Fucini, R.V., Kim, S.M., Masino, R.S., Wespi, R.M., and Miyada, C.G. 1996. Cystic fibrosis mutation detection by hybridization to light-generated DNA probe arrays. Human Mut. 7: 244–255.
Chee, M., Yang, R., Hubbell, E., Berno, A., Huang, X.C., Stern, D., et al. 1996. Accessing genetic information with high-density DNA arrays. Science 274: 610–613.
McIntyre, P.E. 1996. Microfabrication technology for DNA sequencing. Trends Biotech. 14: 69–73.
Sosnowski, R.G., Tu, E., Butler, W.F., O'Connell, J.P., and Heller, M.J. 1997. Rapid determination of single base mismatch mutations in DNA hybrids by direct electric field control. Proc. Wat. Acad. Sci. USA 94: 1119–1123.
Sapolsky, R.J. and Lipshutz, R.J. 1996. Mapping genomic library clones using oligonucleotide arrays. Genomics 33: 445–456.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ramsay, G. DNA chips: State-of-the art. Nat Biotechnol 16, 40–44 (1998). https://doi.org/10.1038/nbt0198-40
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0198-40
This article is cited by
-
ECDEP: identifying essential proteins based on evolutionary community discovery and subcellular localization
BMC Genomics (2024)
-
M2-like polarization of THP-1 monocyte-derived macrophages under chronic iron overload
Annals of Hematology (2020)
-
Innovations in biomedical nanoengineering: nanowell array biosensor
Nano Convergence (2018)
-
Impact of triple-material gate and highly doped source/drain extensions on sensitivity of DNA biosensors
Journal of Computational Electronics (2018)
-
Real-time reliable determination of binding kinetics of DNA hybridization using a multi-channel graphene biosensor
Nature Communications (2017)